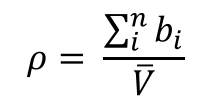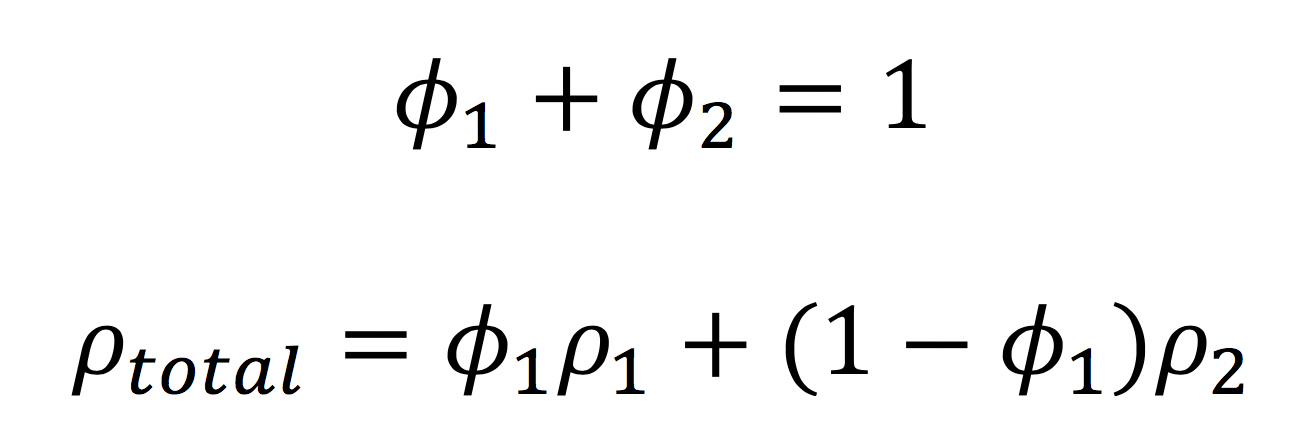| Version 3 (modified by ajj, 7 years ago) (diff) |
|---|
KEMM37 Lab 1B - SANS Data Analysis using SasView
| Table of Contents |
|---|
| 5. Fitting SANS data |
| 5.1 Background Information |
| 5.2 Calculating Scattering Length Density |
| 5.3 Fitting the data |
| Resources |
5. Fitting SANS data
This part of the exercise will use the same real SANS data as the previous exercises - taken from a study of surfactant self assembly in deep eutectic solvents (DES).
TASK 10: Restart SasView
Before starting this part of the exercise, you should have a clean SasView instance. Quit SasView and restart it.
5.1 Background Information
Deep eutectic solvents are a class of ionic liquids formed from a hydrogen bond donor and a halide salt. At a certain mixture ratio, the eutectic mixture, the melting point is significantly depressed to values below room temperature.
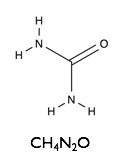
Here we will examine the self-assembly of a surfactant, sodium dodecyl sulfate (SDS) in the deep eutectic solvent formed from a 1:2 molar ratio mixture of choline chloride and urea.
| Sodium Dodecyl Sulfate | Choline Chloride | Urea |
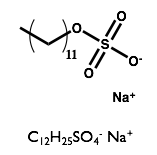 | 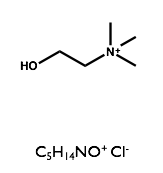 | 
|
TASK 11: If you haven't already done so download the SANS data : SwednessSasViewTutorialData.zip and unzip the file in a known location on your filesystem. Note where you have placed the data.
You should now have a folder called "Subtracted" containing a set of files named as follows:
The data are SANS curves collected on SANS2D at ISIS and D22 at ILL for samples of protonated (normal) SDS in 1:2 d9-choline chloride:d4-urea. This sample was chosen to give maximum contrast and minimum background signal from incoherent scattering. There were 7 samples with 0.2 wt%, 0.5 wt%, 1.0 wt%, 2.0 wt%, 5.0 wt%, 7.5 wt% and 10 wt% of SDS in the DES with the filenames corresponding to each sample given below:
| Surfactant Concentration (wt%) | Data File |
| 0.2 | 0p2hSDS_dChCldUrea_sub.txt |
| 0.5 | 0p5hSDS_dChCldUrea_sub.txt |
| 1.0 | 1hSDS_dChCldUrea_sub.txt |
| 2.0 | 2hSDS_dChCldUrea_sub.txt |
| 5.0 | 5hSDS_dChCldUrea_sub.txt |
| 7.5 | 7p5hSDS_dChCldUrea_sub.txt |
| 10.0 | 10hSDS_dChCldUrea_sub.txt |
All the data files have been processed to 1D scattering curves with the solvent background subtracted to leave only the coherent scattering signal on absolute scale.
Additionally there is a folder called "Not subtracted" containing the data for task 14.
5.2 Calculating Scattering Length Density
The scattering length density is given by
Scattering lengths of relevant elements:
| Element | Scattering Length (fm) |
|---|---|
| C | 6.646 |
| H | -3.739 |
| D | 6.671 |
| N | 9.36 |
| O | 5.803 |
| Cl | 9.577 |
Physical properties of the DES components:
| Component | Chemical Formula | Molecular Volume (Å3) | Density (g/cm3) |
|---|---|---|---|
| d9-Choline Chloride | C5H5D9NOCl | 210.77 | 1.17 |
| d4-Urea | CD4N2O | 75.55 | 1.41 |
TASK 12: Calculate the scattering length density (SLD) of a 1:2 mole ratio mixture of choline chloride and urea.
Use the information in the table above to calculate the SLD. There are multiple ways to do so, including:
- Calculating by hand
- Using a spreadsheet
- Using the SLD calculator built in to SasView (in the Tools menu).
- Using online calculators e.g. https://www.ncnr.nist.gov/resources/activation/
Try calculating by hand and one or more of the other ways and see if you get the same answer!
Remember that in mixtures, we use the volume fraction of each component to calculate the overall scattering length density:
5.3 Fitting the data
TASK 19: Fitting the lowest concentration data.
Select the lowest concentration data only in the data explorer by ensuring only 0p2hSDS_dChCldUrea_sub.txt has a check mark next to it and click “Send to" fitting.
- Select the model for the structure you predicted for this dataset.
- How does it compare to the data?
- Fill in parameters you know and adjust the others to see how close you get to the data.
- Select parameters to fit and run the fit by clicking "Fit" at the bottom of the fitting panel.
- Do you get a good fit?
- Are the parameters you get physically reasonable?
TASK 20: Fitting the other data, starting with the 7.5 wt% data set.
Repeat for other concentrations
- Does the same model fit all data?
- If not, what models do you use?
- What do you observe about the higher concentration data sets?
- What is consistent between datasets? What is different?
TASK 21: Summarising the results
Having fitted all the data sets you can now summarise the results and comment on the trends you observe.
Resources
- NIST SLD calculator https://www.ncnr.nist.gov/resources/activation/
- NIST Scattering Length and Scattering Cross Section Database https://www.ncnr.nist.gov/resources/n-lengths/
Attachments (9)
- SwednessSasViewTutorialData.zip (36.7 KB) - added by ajj 7 years ago.
- kemm37_des.png (16.7 KB) - added by ajj 7 years ago.
- kemm37_sds.png (12.5 KB) - added by ajj 7 years ago.
- kemm37_urea.png (10.9 KB) - added by ajj 7 years ago.
- kemm37_choline.png (10.8 KB) - added by ajj 7 years ago.
- kemm37_loaddata.png (29.5 KB) - added by ajj 7 years ago.
- kemm37_sld.png (11.0 KB) - added by ajj 7 years ago.
- kemm37_volumefraction_sld.png (122.6 KB) - added by ajj 7 years ago.
- kemm37_datafileslist.png (33.8 KB) - added by ajj 7 years ago.
Download all attachments as: .zip